Python3 Read Multiple Files in a Directory
Analyzing Data from Multiple Files
Overview
Teaching: 20 min
Exercises: 0 minQuestions
How tin can I exercise the same operations on many different files?
Objectives
Utilise a library function to get a list of filenames that match a wildcard design.
Write a
forloop to procedure multiple files.
As a terminal piece to processing our inflammation data, nosotros need a style to go a listing of all the files in our information directory whose names start with inflammation- and end with .csv. The following library volition assist us to attain this:
The glob library contains a role, likewise called glob, that finds files and directories whose names match a design. We provide those patterns as strings: the graphic symbol * matches zippo or more characters, while ? matches any one graphic symbol. We can use this to get the names of all the CSV files in the current directory:
print ( glob . glob ( 'inflammation*.csv' )) ['inflammation-05.csv', 'inflammation-11.csv', 'inflammation-12.csv', 'inflammation-08.csv', 'inflammation-03.csv', 'inflammation-06.csv', 'inflammation-09.csv', 'inflammation-07.csv', 'inflammation-10.csv', 'inflammation-02.csv', 'inflammation-04.csv', 'inflammation-01.csv'] Every bit these examples show, glob.glob's issue is a list of file and directory paths in capricious lodge. This ways we tin loop over information technology to exercise something with each filename in turn. In our instance, the "something" we desire to do is generate a set of plots for each file in our inflammation dataset.
If we want to start by analyzing just the beginning three files in alphabetical society, we can utilise the sorted congenital-in function to generate a new sorted list from the glob.glob output:
import glob import numpy import matplotlib.pyplot filenames = sorted ( glob . glob ( 'inflammation*.csv' )) filenames = filenames [ 0 : 3 ] for filename in filenames : print ( filename ) data = numpy . loadtxt ( fname = filename , delimiter = ',' ) fig = matplotlib . pyplot . figure ( figsize = ( 10.0 , iii.0 )) axes1 = fig . add_subplot ( i , 3 , ane ) axes2 = fig . add_subplot ( 1 , 3 , two ) axes3 = fig . add_subplot ( ane , 3 , three ) axes1 . set_ylabel ( 'average' ) axes1 . plot ( numpy . hateful ( data , axis = 0 )) axes2 . set_ylabel ( 'max' ) axes2 . plot ( numpy . max ( data , centrality = 0 )) axes3 . set_ylabel ( 'min' ) axes3 . plot ( numpy . min ( data , centrality = 0 )) fig . tight_layout () matplotlib . pyplot . show () 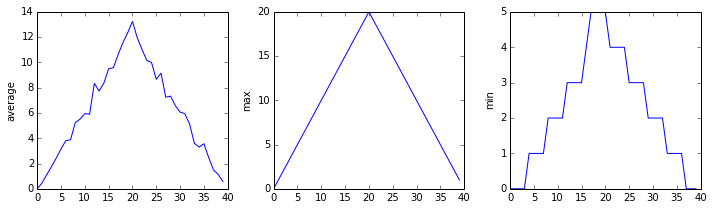
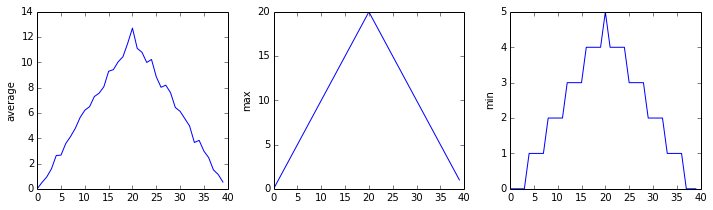
The plots generated for the 2nd clinical trial file look very similar to the plots for the first file: their average plots show similar "noisy" rises and falls; their maxima plots show exactly the same linear rise and fall; and their minima plots prove like staircase structures.
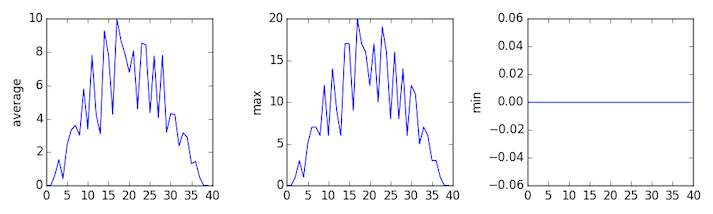
The third dataset shows much noisier average and maxima plots that are far less suspicious than the first two datasets, yet the minima plot shows that the 3rd dataset minima is consistently nada beyond every day of the trial. If we produce a heat map for the third data file we see the post-obit:
We can meet that in that location are zero values sporadically distributed across all patients and days of the clinical trial, suggesting that there were potential issues with information collection throughout the trial. In addition, we tin can see that the concluding patient in the study didn't take any inflammation flare-ups at all throughout the trial, suggesting that they may not even suffer from arthritis!
Plotting Differences
Plot the difference between the boilerplate inflammations reported in the starting time and 2d datasets (stored in
inflammation-01.csvandinflammation-02.csv, correspondingly), i.e., the deviation between the leftmost plots of the commencement two figures.Solution
import glob import numpy import matplotlib.pyplot filenames = sorted ( glob . glob ( 'inflammation*.csv' )) data0 = numpy . loadtxt ( fname = filenames [ 0 ], delimiter = ',' ) data1 = numpy . loadtxt ( fname = filenames [ i ], delimiter = ',' ) fig = matplotlib . pyplot . effigy ( figsize = ( 10.0 , iii.0 )) matplotlib . pyplot . ylabel ( 'Difference in boilerplate' ) matplotlib . pyplot . plot ( numpy . mean ( data0 , axis = 0 ) - numpy . mean ( data1 , axis = 0 )) fig . tight_layout () matplotlib . pyplot . show ()
Generate Composite Statistics
Use each of the files one time to generate a dataset containing values averaged over all patients:
filenames = glob . glob ( 'inflammation*.csv' ) composite_data = numpy . zeros (( 60 , forty )) for filename in filenames : # sum each new file's data into composite_data as information technology's read # # and so divide the composite_data by number of samples composite_data = composite_data / len ( filenames )Then use pyplot to generate average, max, and min for all patients.
Solution
import glob import numpy import matplotlib.pyplot filenames = glob . glob ( 'inflammation*.csv' ) composite_data = numpy . zeros (( 60 , 40 )) for filename in filenames : data = numpy . loadtxt ( fname = filename , delimiter = ',' ) composite_data = composite_data + information composite_data = composite_data / len ( filenames ) fig = matplotlib . pyplot . figure ( figsize = ( 10.0 , 3.0 )) axes1 = fig . add_subplot ( 1 , 3 , 1 ) axes2 = fig . add_subplot ( 1 , 3 , 2 ) axes3 = fig . add_subplot ( one , 3 , 3 ) axes1 . set_ylabel ( 'boilerplate' ) axes1 . plot ( numpy . mean ( composite_data , axis = 0 )) axes2 . set_ylabel ( 'max' ) axes2 . plot ( numpy . max ( composite_data , axis = 0 )) axes3 . set_ylabel ( 'min' ) axes3 . plot ( numpy . min ( composite_data , axis = 0 )) fig . tight_layout () matplotlib . pyplot . show ()
Afterwards spending some fourth dimension investigating the rut map and statistical plots, as well as doing the above exercises to plot differences between datasets and to generate composite patient statistics, nosotros proceeds some insight into the twelve clinical trial datasets.
The datasets appear to fall into two categories:
- seemingly "ideal" datasets that agree excellently with Dr. Maverick'southward claims, simply display suspicious maxima and minima (such every bit
inflammation-01.csvandinflammation-02.csv) - "noisy" datasets that somewhat agree with Dr. Maverick'due south claims, merely show concerning data collection bug such as sporadic missing values and even an unsuitable candidate making it into the clinical trial.
In fact, it appears that all three of the "noisy" datasets (inflammation-03.csv, inflammation-08.csv, and inflammation-11.csv) are identical down to the last value. Armed with this information, we confront Dr. Maverick most the suspicious information and duplicated files.
Dr. Maverick confesses that they fabricated the clinical data after they found out that the initial trial suffered from a number of bug, including unreliable data-recording and poor participant selection. They created fake data to prove their drug worked, and when we asked for more data they tried to generate more fake datasets, too every bit throwing in the original poor-quality dataset a few times to try and make all the trials seem a chip more "realistic".
Congratulations! We've investigated the inflammation data and proven that the datasets accept been synthetically generated.
Just it would be a shame to throw away the synthetic datasets that have taught us so much already, then we'll forgive the imaginary Dr. Bohemian and proceed to use the information to learn how to program.
Key Points
Employ
glob.glob(pattern)to create a listing of files whose names lucifer a pattern.Apply
*in a design to friction match nothing or more characters, and?to match any single character.
Source: https://swcarpentry.github.io/python-novice-inflammation/06-files/index.html
Postar um comentário for "Python3 Read Multiple Files in a Directory"